All Issues
Management changes in rice production alter microbial community
Publication Information
California Agriculture 51(6):33-40. https://doi.org/10.3733/ca.v051n06p33
Published November 01, 1997
PDF | Citation | Permissions
Abstract
Because of an impending ban on burning of residues, more rice growers will be incorporating rice straw in their fields and will depend more than ever on soil microbes to break down the straw and aid rice production the following season. A variety of methods were used to characterize the effects of various rice straw management methods and winter flooding on the microbial community in a typical rice soil. Microbial biomass increased in straw-incorporated plots by the second year, and both flooding and rice straw incorporation caused changes in the relative abundances of specific groups of microorganisms. Although the heavy clay soil and wide variety of soil water contents in this study posed problems for microbial analyses, promising results were obtained with a biochemical characterization of phospholipids of soil microbes. This type of analysis can provide insight into management effects on soil microbial communities.
Full text
More rice growers are incorporating rice straw residues in their fields in anticipation of a ban on burning residues. Scientists are investigating the effects of different straw management methods on the microorganisms that live in the soil.
Agroecosystems depend on the activities of the soil microbial community. Soil microorganisms decompose plant residues, build up soil organic matter, degrade agricultural chemicals, improve soil structure, suppress crop disease and transform nutrients into forms available for plants. When changes in a production system dramatically increase the organic matter added to the soil, the importance of soil microbial activity to productivity becomes more evident. Rice growers have had to adjust to new laws that phase out rice straw burning. Unless a viable market for rice straw is identified, growers will have to incorporate rice straw disposal into their production systems. Another change to rice production management that is being explored is winter flooding of rice fields. The desire to understand the potential impacts of winter flooding is motivated by increasing interest in using fallow rice fields as habitat for migrating birds.
A 75-acre field study of rice straw incorporation methods and winter flooding effects was conducted on a commercial farm near Maxwell in Colusa County.
A primary concern of rice growers who begin to incorporate rice straw in their fields is whether straw decomposes rapidly enough to permit timely planting of the following year's crop. Over the long term, soil organic matter and nutrient cycling changes driven by soil microbial activity and straw inputs may alter fertilizer requirements, disease incidence and pest interactions in the rice production system.
The first step in understanding the potential effects of changes in straw management is to characterize the changes in the soil microbial community resulting from rice straw management and flooding treatments. The microbial biomass is relatively sensitive to perturbations to the soil and, as the most labile fraction of soil organic matter, is an early indicator of changes in soil organic matter in general. Our objective was to characterize the effects of straw management and winter flooding treatments on the size, activity and composition of the soil microbial community.
Measuring soil microorganisms
Soil microbiological studies are limited by the small size of the organisms, the difficulty in extracting them from the soil and the immense diversity of the community. Culture plate methods are highly selective for specific soil microorganisms, possibly representing as few as 1% to 5% of the species present. Those organisms able to form colonies on plates cannot be assumed to be important in overall community functions. Our approach was to use a number of different methods that measured different aspects of the whole soil microbial community. These measures fall into two categories: (1) traditional measures of biomass and activity including fumigation extraction, respiratory activity and direct counts; and (2) new methods that attempt to measure whole community composition or functional abilities and to provide more detail than the traditional measures.
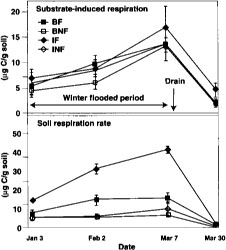
Fig. 1. Substrate induced respiration and basal respiration over time in 1994. BF = burn flooded; BNF = burn not flooded; IF = incorporated flooded; INF = incorporated not flooded. Error bars are standard deviations of four replicate plots.
We attempted to characterize the microbial community as a whole, rather than individual isolates, using two new approaches. One method, substrate utilization analysis (Biolog), measures the decomposition of 95 organic compounds in microtitre plates inoculated with a dilution of the soil. Changes in substrate utilization patterns indicate that those members of the microbial community able to grow on the plates have changed with respect to their metabolic capacity. This pattern can be used as a fingerprint of the culturable microbial community and can be compared among treatments.
The second method, phospholipid fatty acid (PLFA) analysis, differs substantially from substrate utilization analysis because PLFA requires neither growth nor metabolism of the soil community in vitro. Fatty acids, which make up living cell membranes, are extracted directly from soil and identified with gas chromatography. This method provides a fingerprint of the microbial community that reflects the species composition of the community, rather than its metabolic capabilities. In addition, the total amount of PLFA can be used to estimate the biomass of microorganisms, and individual PLFAs can be used as biomarkers to quantify specific groups of microorganisms, such as fungi, bacteria, actinomycetes, anaerobes and methane oxidizers.
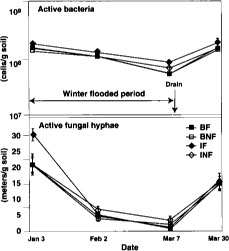
Fig. 2. Direct counts of active bacteria and fungal hyphae over time in 1994. BF = burn flooded; BNF = burn not flooded; IF = incorporated flooded; INF = incorporated not flooded. Error bars are standard deviations of four replicate plots.
It was particularly challenging to study microbial ecology in this rice production system because soil water content ranged widely in the plots and the soil was a heavy clay. A number of different methods were used to characterize the microbial community in this rice soil. Because this broad range of methods — including the two newer methods that characterize the functional and compositional changes in the soil microbial community — have rarely been used together in a replicated field trial, their strengths and limitations in measuring soil microorganisms are discussed.
A 75-acre field study of rice straw incorporation methods and winter flooding effects was initiated in October 1993 on a commercial farm near Maxwell in Colusa County. It was a randomized complete-block experiment with split plots; two water treatments were used on the main plots (winter flood or no flood), and four rice straw management methods were used on the split plots (straw burned, removed, rolled or incorporated). All treatments were replicated four times.
In the first year, highly contrasting treatment plots (straw burned and straw incorporated plots, both flooded and not flooded) were sampled four times. Representative surface soil samples (0-6 inches) were taken by pooling 20 cores from a 328-foot (100 m) transect in each plot in early winter (Jan. 3); midwinter (Feb. 2); predrain (March 7); and 1-month postdrain (March 30, 1994). In 1995 samples were taken from all treatment plots twice: predrain (Feb. 27) and postdrain (April 10).
As a field index of microbial activity, mesh bags filled with chopped rice straw were buried in treatment plots on Feb. 2, 1994, along the same transect used for soil sampling. Bags were sampled Feb. 17, March 7, March 18 and March 29, 1994.
Total microbial biomass carbon and nitrogen were determined in 1994 samples by fumigation extraction. Active microbial biomass was measured in three ways in the 1994 samples: (1) substrate-induced respiration (glucose added) measured as CO2 production; (2) direct counts of fluorescein diacetate-active (FDA) bacteria and fungal hyphae length; and (3) total phospholipid fatty acid analysis (PLFA).
In 1995 samples, active microbial biomass was measured by total PLFA and direct counts of FDA-active bacteria and fungi. Microbial activity was measured by determining basal respiration (no glucose substrate added) as CO2 production in 1994 samples. In 1995 basal respiration was measured in samples at field moisture (that is, unmixed and without addition of water).
To analyze changes in microbial community function, we used a new method based on sole-carbon-source utilization. Carbon-source utilization patterns were determined on Biolog GN microplates inoculated with a soil dilution and incubated.
PLFA analysis was conducted by extracting triplicate subsamples with a one-phase extraction mixture containing chloroformmethanol: phosphate buffer. Phospholipids were separated from neutral lipids and glycolipids, then subjected to mild alkaline methanolysis. Samples were analyzed by gas chromatography.
Analysis of variance was used to test for significant effects of treatments on traditional measures of microbial biomass and activity. Multivariate statistical analysis techniques used in vegetation community ecology, redundancy analysis (RDA) and canonical correspondence analysis (CCA) were used to determine the impact of treatments on carbon-source utilization patterns and PLFA profiles. These analyses are very similar to principal components analysis in that similar samples are grouped together and dissimilar samples are separated along artificial axes created by the analysis.
Fig. 3. Soil respiration rates in February 1995, before draining of flooded plots, and April 1995, 1-month postdraining. Error bars are standard deviations of four replicate plots.
Fig. 4. Straw decomposition in mesh bags buried in straw-burned and straw-incorporated, flooded and not flooded plots in 1994. BF = burn flooded; BNF = burn not flooded; IF = incorporated flooded; INF = incorporated not flooded. Error bars are standard deviations of four replicate plots.
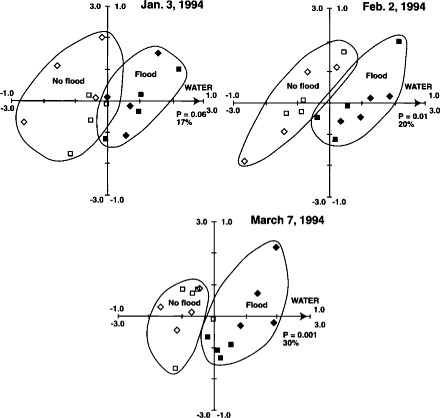
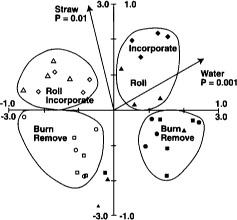
Fig. 5. Canonical correspondence analysis ordination biplot of Biolog substrate-utilization pattern treatment plot scores and significant environmental variables for Jan. 3, Feb. 2 and March 7 sample dates from Maxwell site. ⊡ = straw incorporated, no flooding; ● = straw incorporated, winter flooding; ▪ = straw burned, no flooding; ▴ = straw burned, winter flooding. The 1.0 scale refers to environmental variables; the 3.0 scale refers to treatment plot scores. Reprinted by permission from the American Society for Microbiology.Samples are plotted based on weighted averages of Biolog well scores (or relative abundance of PLFAs in later figures). Arrows show direction of high carbon inputs, soil water contents (flooding) or date. Sample plots close together are most similar; plots farthest apart are most different. Length of axes representing environmental variables indicates importance variables had in explaining variation in data. Significance levels (P values) for environmental variables based on the Monte Carlo permutation test.
With RDA and CCA, however, environmental variables — such as carbon inputs, soil water or straw management regime — are used to constrain the created axes in such a way that the relationship between the multivariate data and the known, measured environmental variables is illustrated and quantified.
The Monte Carlo permutation test was used to test the statistical significance of the variables. Polygons have been drawn around like treatment plots on the figures to facilitate interpretation.
Changes in soil microbiology
Microbial biomass
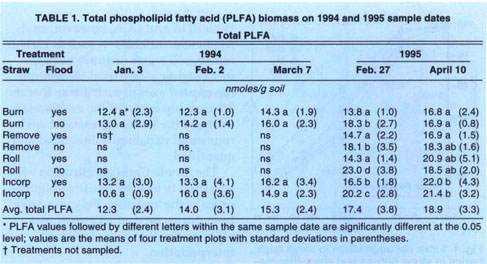
Microbial biomass did not differ with management practices in 1994. Fumigation extraction carbon ranged from 76 μC/g soil to 114 μgC/g soil over the 1995 sample season, but treatments did not differ, nor did the treatments differ in fumigation extraction nitrogen, which ranged from 3 μgN/g soil to 6.5 μgN/g soil. Measures of active microbial biomass — substrate, induced respiration (fig. 1), direct counts (fig. 2) and total PLFA (table 1) — were also similar in all treatments, except that small in creases in bacteria direct counts were found in incorporated flooded plots. Active fungi declined over the winter in 1994 and were lower in flooded plots by March (fig. 2). This was in accord with the fact that fungi are less competitive than bacteria in wet soils. In 1995 microbial biomass as measured by total PLFA increased in straw-incorporated plots compared to all other treatments and was lower in winter flooded than unflooded plots during the flooded period (table 1). Microbial biomass showed smaller increases in straw-rolled than in straw-incorporated plots, which was probably due to less contact between straw and soil in the rolled plots. Active bacteria counts showed no difference in 1995, however, and active hyphae counts were variable and not significantly different.
Basal respiration
Measured respiration was high and increased over time in flooded plots (fig. 1), decreasing after the drain in all plots. This was probably due to a buildup in flooded plots of anaerobic decomposition products that were readily metabolized when samples were aerated during soil sampling and analysis. This effect was less apparent when respiration was measured in 1995, when the methodology resulted in less disturbance of the samples (fig. 3). One month after draining of flooded plots in 1995, basal respiration was lowest in straw-burned and -removed plots, and highest in straw-incorporated plots (fig. 3). Still, differences were small at this time, probably indicating that readily available organic matter had substantially decomposed in straw-incorporated plots.
Straw bag decomposition
Straw in mesh bags buried in treatment plots in 1994 decomposed more slowly in not-flooded plots in which straw was burned than in the other plots (fig. 4). Across all straw treatments, winter flooding enhanced rates of decomposition. This was unexpected because decomposition was assumed to be slower under the more anaerobic conditions present in the flooded soils. Flooding may have had a positive impact on decomposition rates because of increased temperature buffering or increased contact between microorganisms and the straw. It is also possible that these soils, which are adapted to periods of flooding, have a higher potential for anaerobic decomposition than previously assumed.
Substrate utilization patterns (Biolog)
Substrate utilization patterns as a fingerprint of the soil bacterial community were reproducible and sensitive to both straw incorporation and flooding in the first year of treatments (fig. 5). Biolog patterns were similar in flooded and not-flooded plots in the second year, however, probably because the very wet winter of 1995 diminished the difference between the treatments. Substrate utilization patterns were relatively insensitive to temporal changes between winter and spring.
Phospholipid fatty acid analysis
PLFA profiles provided replicable fingerprints of the soil microbial community. Changes in the community that were strongly correlated with flooding increased over the duration of flooding (fig. 6). This is evident by the increased significance (P = 0.06 to P = 0.001) of the soil water variable from Jan. 3 to March 7, 1994, and by the increased amount of variation explained by the axis constrained by the soil water gradient (17% in January to 30% in March). Straw management caused measurable changes in PLFA profiles in the second but not the first year of treatments; thus the straw-incorporated and straw-rolled plots separate completely from the burned and removed plots in March 1995 (fig. 7). Flooding was also highly significant in March 1995 (during the winter flooded period), consistent with the 1994 results. After the drain in April 1995, straw-incorporated plots were found to be significantly different from straw-rolled plots (fig. 8), whereas straw-rolled plots were not different from straw-removed plots. Temporal shifts in microbial communities were greater than differences associated with treatments, yet differences specific to straw management and flooding were still clearly distinguishable. This was in contrast with Biolog results, in which changes over time were much smaller than changes associated with management.
Strengths and limits of measures
The multitude of approaches available for describing microbial communities often makes it difficult to decide which methods are appropriate for a particular study. Several methods were used the first year of this study because of the uncertainty about which were feasible to use at this field site, which had heavy clay soil and a wide range of soil water contents.
Uncertainties in the interpretation of results from several measures resulted from the wide range in soil water content. Fumigation extraction for estimating biomass depends on diffusion of chloroform into soil, and thus extraction efficiency may have varied among treatment plot types owing to different amounts of water. The heavy clay soil also impedes diffusion so it would be difficult to compare these results of fumigation extraction with those from a lighter-textured soil.
Fig. 6. Redundancy analysis ordination biplots of PLFA profiles, with the first ordination axis representing soil water contents for Jan. 3, Feb. 2 and March 7, 1994. ▪ = straw burned; ♦ = straw incorporated; open symbols = not flooded. Axis scale and biplot interpretation as described in Fig. 5. Reprinted by permission from Springer-Verlag.
Fig. 7. Redundancy analysis ordination biplot of PLFA profiles and significant environmental variables from Feb. 27, 1995. ▪ = straw burned; ▪ = straw removed; ● = straw rolled; ▴ = straw incorporated; and open symbol = not flooded. Axis scale and biplot interpretation as described in Fig. 5. Reprinted by permission from Springer-Verlag.
Fig. 8. Redundancy analysis ordination biplot of PLFA profiles, centroids of straw management regime and the soil water content axis from April 10, 1995. ▪ = straw burned; ● = straw removed; ▴ = straw rolled; ♦ = straw incorporated; open symbols = not flooded. Reprinted by permission from Springer-Verlag.Management variables such as straw incorporation, rolling, removal or burning can also be included as constraining environmental variables in ordination. They are represented by points on the biplot (centroids) that fall in the middle of points associated with that variable.
Measures of respiration were problematic in samples from flooded treatments also because sampling and analysis introduced oxygen into the samples. High rates of respiration were found in samples from flooded plots (fig. 1). This was probably caused by the rapid aerobic decomposition, during the analysis, of readily available carbon substrates, such as organic acids, built up under anaerobic conditions. Standing water varying from none to several inches would also cause variation in a field measure of respiration activity.
In 1995, after the soil was drained and the different plots had similar water contents, the laboratory measure of CO2 release was sensitive to differences in straw placement, for example, in incorporated versus straw-rolled plots (fig. 3). As a measure of readily available substrate in similarly aerated soil, it is very useful. Because the measure is sensitive to dilution factors, direct counts of active bacteria could also be affected by flooding.
Treatments with varying amounts of straw incorporated were problematic for the direct counts of active bacteria and fungi, which were confounded by significant clumping of microorganisms on pieces of decomposing straw. These clumps, which were most abundant in plots where straw was rolled or incorporated, contained hundreds of individual bacteria, too numerous to count. This measure is not reliable for contrasting microbial populations in soils receiving different amounts of incorporated organic matter.
Substrate utilization patterns were highly sensitive, particularly to straw inputs. However, PLFA analysis detected changes in the microbial community structure that were not evident with the Biolog plate measure of function. It is clear that substrate utilization cannot be assumed to reflect differences in microbial community composition. The substrate utilization method probably fails to provide conditions appropriate for many organisms, and not all members of a community are metabolically active. Whether the subset of the community detected by Biolog substrate utilization is indeed important to the rice soils cannot be determined in this study.
PLFA profiles provided sensitive fingerprints of microbial communities, and these profiles provided more information about the community than did many of the other measures used. The extraction procedure, involving harsh solvents, is likely to retrieve more cells than the direct count extraction procedure and is not affected by soil moisture, as is fumigation extraction. Therefore this method is the least likely of the methods used to be biased by the nature of the treatments and soil type encountered in this study.
Community composition shifts
The relative amounts of specific groups of fatty acids were changed by straw treatments and winter flooding. In an attempt to relate those fatty acids that changed to specific taxonomic or functional groups of microorganisms, we compared the field data to specific peaks reported to be associated with certain microbial groups. In several cases, existing data were useful in interpreting the results (table 2). For example, fungal populations were lower under flooded conditions and higher in plots with straw incorporation after 2 years. Anaerobic bacteria were more abundant in flooded plots. A number of the PLFAs that responded to management or season are ones with which no specific taxonomic or functional association has yet been made.
When compared with PLFA profiles from the Sustainable Agriculture Farming Systems (SAFS) project at UC Davis, it was evident that all treatments in the rice soil had smaller proportions of fungi and actinomycetes than the SAFS soil, and larger proportions of bacterial fatty acids that have been previously associated with species selected under oxygen limitations. The same set of fatty acids were enriched in both SAFS and rice soils that received high amounts of organic matter. This suggested that similar groups of microorganisms respond to organic inputs in two soils that differed substantially in soil type and agricultural management.
Our study is one of few to successfully apply PLFA analysis in a replicated agricultural field trial. Numerous questions concerning the interpretation of the changes that occur in the fingerprints remain, particularly with respect to which particular organisms or functional groups are sensitive to management. Many of the PLFAs proposed to be biomarkers for specific microbial groups (for example, methanotrophs) have their basis in laboratory studies of only a few isolated members of a particular group. Therefore the biomarkers may not be representative of all members of that group occurring in nature, nor are they necessarily exclusive to the group that they are supposed to represent. Future research is needed to relate those PLFAs consistently enriched or reduced by certain management practices to specific groups of microorganisms. Nevertheless, the essentially unbiased nature of the assay — that is, it measures a specific type of molecule from all organisms present in the soil — makes it the most promising new technique for whole soil microbial community studies readily available at present. Some of the newly emerging DNA- or RNA-based methods for characterizing whole communities may soon provide other alternatives.
Summary of changes
We found no change in the overall size of the microbial biomass in the first year of straw treatments and winter flooding. However, changes did occur in the composition of the microbial community and possibly in its functional abilities because winter flooding treatments. By the second year, changes in the microbial community composition from both winter flooding and straw incorporation were evident, and an overall increase in the microbial biomass had resulted from straw inputs. These data indicate that the long-term result of on-site straw disposal, whether incorporated or rolled, will be an increase in the size of the active microbial community. The microbial biomass represents an actively cycling pool of nutrients, and changes in this pool size have been associated with altered nutrient dynamics in other systems. Linking the changes observed in the microbial community of rice soils to nutrient dynamics during the rice-growing season should be a research priority. Long-term changes in decomposition rates in this soil resulting from higher microbial biomass may also occur.
Contrary to expectations, winter flooding caused somewhat faster decomposition of incorporated straw. Therefore providing overwintering habitat for waterfowl is apparently not in conflict with on-farm straw disposal. At the International Rice Research Institute in the Philippines, however, researchers have found that when soil is maintained in a flooded condition for much of the year, altered decomposition pathways change the soil organic matter. These changes in the quality of the organic matter have been implicated in sequestration of nitrogen and yield declines. We identified many specific changes in PLFA profiles of the microbial community resulting from winter flooding. The detailed look at the active microbial community provided by PLFA analysis may provide an opportunity to look at processes that may, in the long term, result in important changes in soil organic matter quality.
The variety of methods used in this study together provide a robust foundation of baseline information about the microbial community in this rice soil. Early impacts of rice straw management and winter flooding treatments have been characterized based on both traditional and promising new methods. The observed changes in the microbial community can be associated with important aspects of the system, such as rice straw decomposition rates, disease suppression, methane production and denitrification rates, to provide an understanding of the interactions between management, soil microbes and important agroecosystem functions. Such analyses can provide a basis for an understanding of the manageable aspects of the soil microbial activity.